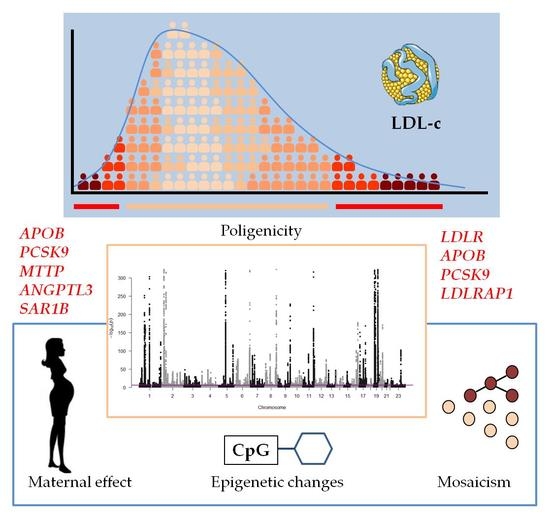
Genetic Determinants of Plasma Low-Density Lipoprotein Cholesterol Levels: Monogenicity, Polygenicity, and "Missing" Heritability
Source : https://www.mdpi.com/2227-9059/9/11/1728
Changes in plasma low-density lipoprotein cholesterol (LDL-c) levels relate to a high risk of developing some common and complex diseases. LDL-c, as a quantitative trait, is multifactorial and depends on both genetic and environmental factors. In the pregenomic age, targeted genes were used to detect genetic factors in both hyper- and hypolipidemias, but this approach only explained extreme cases in the population distribution.
This paper discusses other possibilities for unexplained dyslipidemias associated with LDL-c, such as mosaicism, maternal effect, and induced epigenetic changes. Future studies should consider gene–gene and gene–environment interactions and the development of integrated information about disease-driving networks, including phenotypes, genotypes, transcription, proteins, metabolites, and epigenetics.





• Source: Biomedicines
• Conclusion/Relevance: “This paper discusses other possibilities for unexplained dyslipidemias associated with LDL-c, such as mosaicism, maternal effect, and induced epigenetic changes. Future studies should consider gene–gene and gene–environment interactions and the development of integrated information about disease-driving networks, including phenotypes, genotypes, transcription, proteins, metabolites, and epigenetics.”
• GWASs have for the most part supplanted previous family linkage studies that relied genomic markers (i.e., microsatellites) to detect candidate regions. Linkage studies assess for chromosomal segments that cosegregate with a given trait. They rely on extended pedigrees and analyzing various generations of affected families, and assume limited recombination events and low recurrent mutation rates.
• Although GWAS regions are broad, they demonstrate highly penetrant phenotypes, which are indicative of rare diseases.
• “However, GWASs look for traits associated with a given chromosomal segment, using unrelated individuals and, thus, considering many generations of recombination,” wrote the authors. “Unlike microsatellites, which have higher mutation rates, most variants used in GWASs are single-nucleotide variants (SNVs) that can be considered identical by descent. This works well for low penetrant phenotypes, typical of most common diseases.”
• The concept of polygenic origin refers to SNVs linked to plasma LDL-c.
• LDL-c raising alleles and polygenic risk scores (PRSs) determine the contribution of the accumulation of small-effect variants to plasma LDL-c.